89-000-736
heat shock transcription factor 4, Mouse, Clone: 3G3, Abnova™
Manufacturer: Abnova Corporation
Select a Size
| Pack Size | SKU | Availability | Price |
|---|---|---|---|
| Each of 1 | 89-000-736-Each-of-1 | In Stock | ₹ 42,898.00 |
89-000-736 - Each of 1
In Stock
Quantity
1
Base Price: ₹ 42,898.00
GST (18%): ₹ 7,721.64
Total Price: ₹ 50,619.64
Antigen
heat shock transcription factor 4
Classification
Monoclonal
Conjugate
Unconjugated
Formulation
PBS with no preservative; pH 7.4
Gene Accession No.
NM_001538
Gene Symbols
HSF4
Immunogen
HSF4 (NP_001529, 121 a.a. ∼ 220 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa.
Quantity
100 μg
Research Discipline
Transcription Regulation
Primary or Secondary
Primary
Target Species
Human
Product Type
Antibody
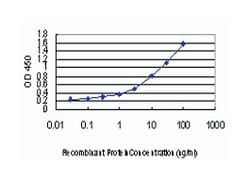
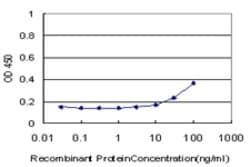
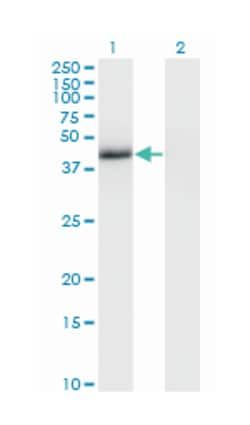
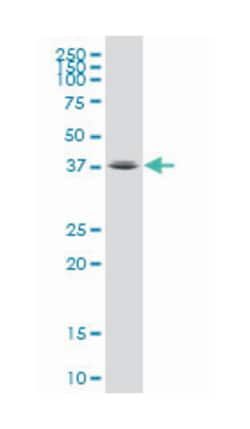
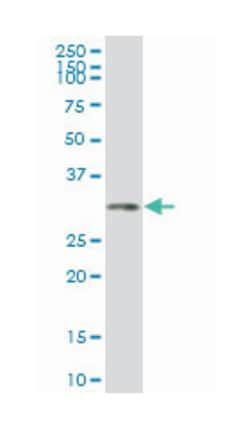
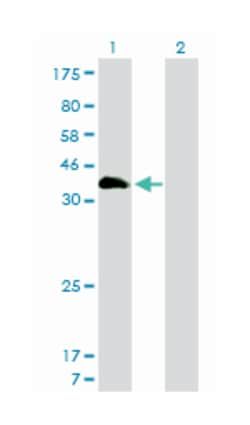
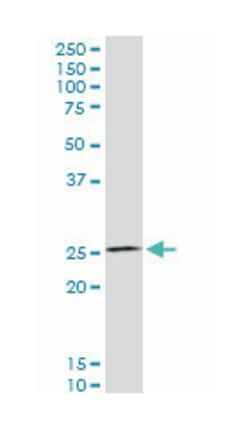
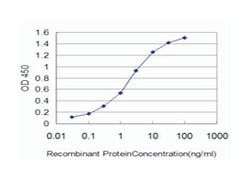
Applications
ELISA, Western Blot
Clone
3G3
Description
Mouse monoclonal antibody raised against a partial recombinant HSF4.
Gene
HSF4
Gene Alias
CTM
Host Species
Mouse
Purification Method
Affinity Purified
Regulatory Status
RUO
Whole Molecule
Yes
Gene ID (Entrez)
3299
Content And Storage
Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing.
Isotype
IgG2a κ
Description
- Heat-shock transcription factors (HSFs) activate heat-shock response genes under conditions of heat or other stresses
- HSF4 lacks the carboxyl-terminal hydrophobic repeat which is shared among all vertebrate HSFs and has been suggested to be involved in the negative regulation of DNA binding activity
- Two alternatively spliced transcripts encoding distinct isoforms and possessing different transcriptional activity have been described
- [provided by RefSeq Sequence: KVPALRGDDGRWRPEDLGRLLGEVQALRGVQESTEARLRELRQQNEILWREVVTLRQSHGQQHRVIGKLIQCLFGPLQAGPSNAGGKRKLSLMLDEGSSC