89-000-924
RAD1 homolog (S. pombe), Mouse, Clone: 1G2, Abnova™
Manufacturer: Abnova Corporation
Select a Size
| Pack Size | SKU | Availability | Price |
|---|---|---|---|
| Each of 1 | 89-000-924-Each-of-1 | In Stock | ₹ 44,411.00 |
89-000-924 - Each of 1
In Stock
Quantity
1
Base Price: ₹ 44,411.00
GST (18%): ₹ 7,993.98
Total Price: ₹ 52,404.98
Antigen
RAD1 homolog (S. pombe)
Classification
Monoclonal
Conjugate
Unconjugated
Formulation
PBS with no preservative; pH 7.4
Gene Accession No.
BC006837
Gene Symbols
RAD1
Immunogen
RAD1 (AAH06837, 1 a.a. ∼ 90 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa.
Quantity
100 μg
Whole Molecule
Yes
Gene ID (Entrez)
5810
Content And Storage
Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing.
Isotype
IgG3 κ
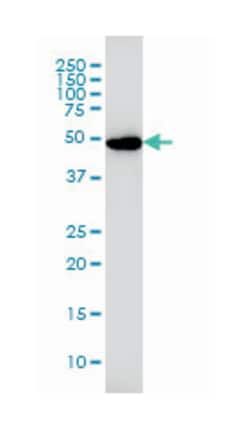
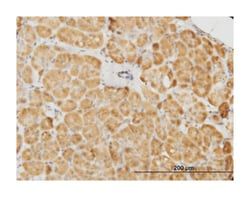
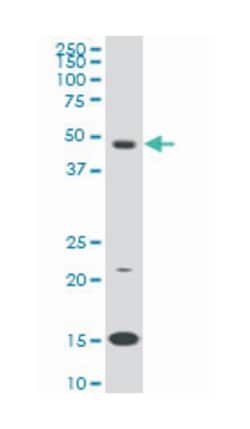
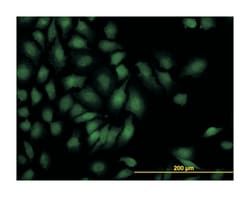
Applications
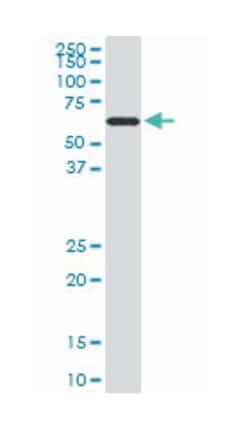
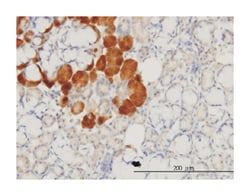
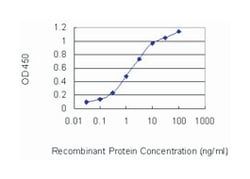
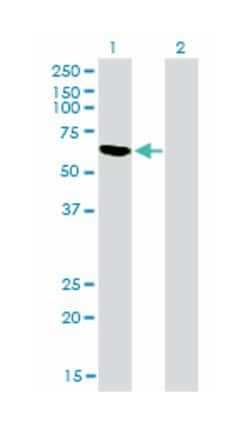
ELISA, Immunohistochemistry (PFA fixed), KnockDown, Western Blot
Clone
1G2
Description
Mouse monoclonal antibody raised against a partial recombinant RAD1.
Gene
RAD1
Gene Alias
HRAD1/REC1
Host Species
Mouse
Purification Method
Affinity Purified
Regulatory Status
RUO
Primary or Secondary
Primary
Target Species
Human
Product Type
Antibody
Description
- This gene encodes a component of a heterotrimeric cell cycle checkpoint complex, known as the 9-1-1 complex, that is activated to stop cell cycle progression in response to DNA damage or incomplete DNA replication
- The 9-1-1 complex is recruited by RAD17 to affected sites where it may attract specialized DNA polymerases and other DNA repair effectors
- Alternatively spliced transcript variants of this gene have been described
- [provided by RefSeq Sequence: MPLLTQQIQDEDDQYSLVASLDNVRNLSTILKAIHFREHATCFATKNGIKVTVENAKCVQANAFIQAGIFQEFKVQEESVTFRINLTVLL