89-004-954
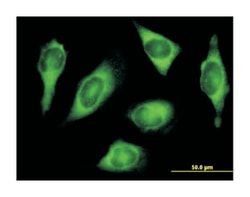
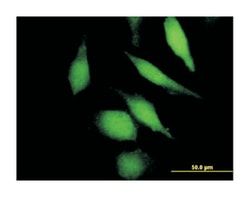
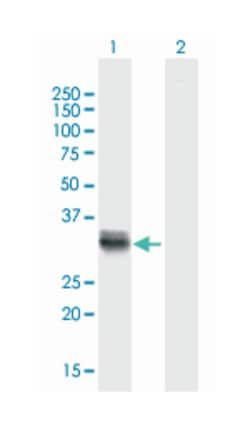
ATPase, H+/K+ exchanging, beta polypeptide, Mouse, Polyclonal Antibody, Abnova™
Manufacturer: Abnova Corporation
Select a Size
| Pack Size | SKU | Availability | Price |
|---|---|---|---|
| Each of 1 | 89-004-954-Each-of-1 | In Stock | ₹ 42,898.00 |
89-004-954 - Each of 1
In Stock
Quantity
1
Base Price: ₹ 42,898.00
GST (18%): ₹ 7,721.64
Total Price: ₹ 50,619.64
Antigen
ATPase, H+/K+ exchanging, beta polypeptide
Classification
Polyclonal
Description
Mouse polyclonal antibody raised against a full-length human ATP4B protein.
Gene
ATP4B
Gene Alias
ATP6B
Host Species
Mouse
Quantity
50 μL
Whole Molecule
Yes
Gene ID (Entrez)
496
Content And Storage
Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing.
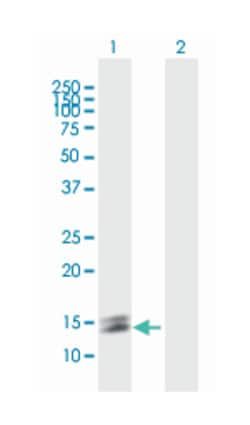
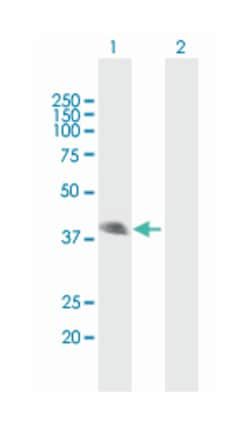
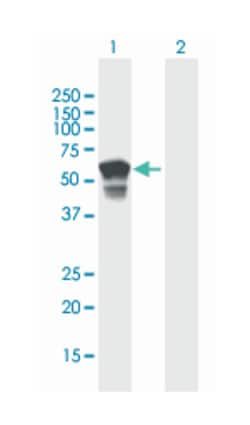
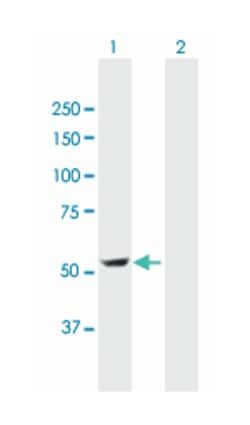
Applications
Western Blot
Conjugate
Unconjugated
Formulation
No additive
Gene Accession No.
NM_000705.2
Gene Symbols
ATP4B
Immunogen
ATP4B (NP_000696.1, 1 a.a. ∼ 291 a.a) full-length human protein.
Regulatory Status
RUO
Primary or Secondary
Primary
Target Species
Human
Form
Serum
Description
- The protein encoded by this gene belongs to a family of P-type cation-transporting ATPases
- The gastric H+, K+-ATPase is a heterodimer consisting of a high molecular weight catalytic alpha subunit and a smaller but heavily glycosylated beta subunit
- This enzyme is a proton pump that catalyzes the hydrolysis of ATP coupled with the exchange of H(+) and K(+) ions across the plasma membrane
- It is also responsible for gastric acid secretion
- This gene encodes the beta subunit of the gastric H+, K+-ATPase
- [provided by RefSeq] Sequence: MAALQEKKTCGQRMEEFQRYCWNPDTGQMLGRTLSRWVWISLYYVAFYVVMTGLFALCLYVLMQTVDPYTPDYQDQLRSPGVTLRPDVYGEKGLEIVYNVSDNRTWADLTQTLHAFLAGYSPAAQEDSINCTSEQYFFQESFRAPNHTKFSCKFTADMLQNCSGLADPNFGFEEGKPCFIIKMNRIVKFLPSNGSAPRVDCAFLDQPRELGQPLQVKYYPPNGTFSLHYFPYYGKKAQPHYSNPLVAAKLLNIPRNAEVAIVCKVMAEHVTFNNPHDPYEGKVEFKLKIEK